Cell membranes are fascinating supramolecular aggregates that not only form a barrier between different compartments, or organells, of cells but also harbor many chemical reactions essential to the existence and functioning of a cell. For example, the plasma membrane serves as a barrier to prevent the contents of a cell from escaping and mixing with the surrounding medium. At the same time a cell’s plasma membrane must enable the passage of critical nutrients into the cell and the passage of waste products out. Membranes must also be flexible to enable cells to change shape and they must also have malleable topologies such that a cell can grow and divide into two separate parts, each of which has a completely closed contiguous membrane.
Membranes in living biological systems have managed to balance these multiple demands by exploiting the special amphiphilic physical properties of the molecules that make them up. The most common membrane constituents are lipid molecules with two physically separated subdomains, usually an elongated hydrophobic domain made up of fatty acid tails, associated with a hydrophilic head group. The most abundant lipids in cell membranes are the phospholipids, molecules in which the hydrophilic head is linked to the rest of the lipid through a phosphate group. The hydrophilic head groups of lipids dissolve readily in water because they contain polar groups that are easily incorporated in the hydrogen-bonding network of the surrounding water.
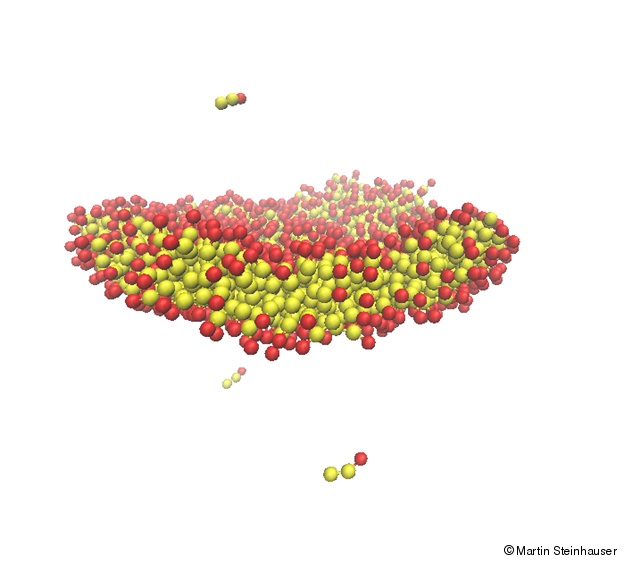
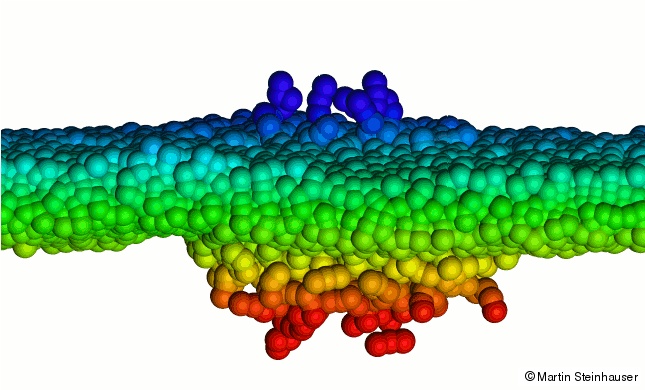
In contrast, the hydrophobic hydrocarbon tails are uncharged and nonpolar and thus try to aggregate in energetically and entropically favorable structures that minimize their contact with surrounding water molecules. The nearly cylindrical shape of most membrane lipids makes the bilayer the most common geometrical organization for spontaneous self-assembly of lipid molecules in aqueous environment (see top figure). Many questions pertaining to cell membrane dynamics remain unanswered until today and many processes such as lateral segregation or transversal asymmetry are known to occur in membranes on vastly different spatiotemporal scales. It is therefore not yet possible to visualize the re-formation of the lipids in a fusion pore experimentally. It is precisely at the spatial and temporal limits where instruments for direct observation fail that the theoretical membrane concept becomes fuzzy and unclear.
We present here a MD study of the clustering of lipid molecules, which start from random initial configurations and finally form stable bilayer membranes. We use an implicit solvent model in my code MD-Cube and investigate the successive steps in membrane formation with respect to the degree of order that is introduced to the system by the molecular self-assembly. The dynamic processes of lipid clustering are investigated using Langevin dynamics. We find that the membrane self-assembly undergoes several stages of clustering that subsequently minimize the free energy of the system, and finally lead to a stable, fluid-like bilayer structure where the lipid molecules exhibit strong fluctuations. The different stages of clustering can also be seen when monitoring the total potential energy of the system.
Selected publications
Destruction of cancer cells by laser-induced shockwaves: recent developments in experimental treatments and multiscale computer simulations
M.O. Steinhauser, M. Schmidt
SoftMatter, 2014, 10, 4778–4788
Computational Multiscale Modeling of Fluids and Solids
M.O. Steinhauser
Springer, Berlin, Heidelberg, 2017
Multiscale modeling, coarse-graining and shock wave computer simulations in materials science
M.O. Steinhauser
AIMS Materials Science, 4(6): 1319-1357